Abstract
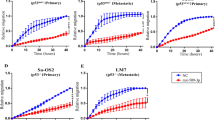
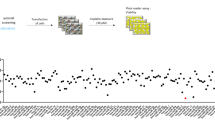
In this study, high-throughput microRNA (miRNA) expression analysis revealed that the expression of miR-140 was associated with chemosensitivity in osteosarcoma tumor xenografts. Tumor cells ectopically transfected with miR-140 were more resistant to methotrexate and 5-fluorouracil (5-FU). Overexpression of miR-140 inhibited cell proliferation in both osteosarcoma U-2 OS (wt-p53) and colon cancer HCT 116 (wt-p53) cell lines, but less so in osteosarcoma MG63 (mut-p53) and colon cancer HCT 116 (null-p53) cell lines. miR-140 induced p53 and p21 expression accompanied with G1 and G2 phase arrest only in cell lines containing wild type of p53. Histone deacetylase 4 (HDAC4) was confirmed to be one of the important targets of miR-140. The expression of endogenous miR-140 was significantly elevated in CD133+hiCD44+hi colon cancer stem-like cells that exhibit slow proliferating rate and chemoresistance. Blocking endogenous miR-140 by locked nucleic acid-modified anti-miR partially sensitized resistant colon cancer stem-like cells to 5-FU treatment. Taken together, our findings indicate that miR-140 is involved in the chemoresistance by reduced cell proliferation through G1 and G2 phase arrest mediated in part through the suppression of HDAC4. miR-140 may be a candidate target to develop novel therapeutic strategy to overcome drug resistance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Alvarez-Garcia I, Miska EA . (2005). MicroRNA functions in animal development and human disease. Development 132: 4653–4662.
Ambros V, Lee RC . (2004). Identification of microRNAs and other tiny noncoding RNAs by cDNA cloning. Methods Mol Biol 265: 131–158.
Bartel DP . (2004). MicroRNAs: genomics, biogenesis, mechanism and function. Cell 116: 281–297.
Botchkina IL, Rowehl RA, Rivadeneira DE, Karpeh Jr MS, Crawford H, Dufour A et al. (2009). Phenotypic subpopulations of metastatic colon cancer stem cells: genomic analysis. Cancer Genomics Proteomics 6: 19–29.
Braun CJ, Zhang X, Savelyeva I, Wolff S, Moll UM, Schepeler T et al. (2008). p53-responsive microRNAs 192 and 215 are capable of inducing cell cycle arrest. Cancer Res 68: 10094–10104.
Bruheim S, Bruland OS, Breistol K, Maelandsmo GM, Fodstad O . (2004). Human osteosarcoma xenografts and their sensitivity to chemotherapy. Pathol Oncol Res 10: 133–141.
Bunz F, Hwang PM, Torrance C, Waldman T, Zhang Y, Dillehay L et al. (1999). Disruption of p53 in human cancer cells alters the responses to therapeutic agents. J Clin Invest 104: 263–269.
Calin G, Croce CM . (2006). MicroRNA signatures in human cancers. Nat Rev Cancer 6: 857–866.
Chandar N, Billig B, McMaster J, Novak J . (1992). Inactivation of p53 gene in human and murine osteosarcoma cells. Br J Cancer 65: 208–214.
Dalerba P, Dylla SJ, Park IK, Liu R, Wang X, Cho RW et al. (2007). Phenotypic characterization of human colorectal cancer stem cells. Proc Natl Acad Sci USA 104: 10158–10163.
Dressler LG, Seamer LC, Owens MA, Clark GM, McGuire WL . (1988). DNA flow cytometry and prognostic factors in 1331 frozen breast cancer specimens. Cancer 61: 420–427.
Du L, Wang H, He L, Zhang J, Ni B, Wang X et al. (2008). CD44 is of functional importance for colorectal cancer stem cells. Clin Cancer Res 14: 6751–6760.
Eberharter A, Becker PB . (2002). Histone acetylation: a switch between repressive and permissive chromatin. Second in review series on chromatin dynamics. EMBO Rep 3: 224–229.
Finnin MS, Donigian JR, Cohen A, Richon VM, Rifkind RA, Marks PA et al. (1999). Structures of a histone deacetylase homologue bound to the TSA and SAHA inhibitors. Nature 401: 188–193.
Georges SA, Biery MC, Kim SY, Schelter JM, Guo J, Chang AN et al. (2008). Coordinated regulation of cell cycle transcripts by p53-inducible microRNAs, miR-192 and miR-215. Cancer Res 68: 10105–10112.
Grozinger C, Schreiber SL . (2002). Deacetylase enzymes: biological functions and the use of small-molecule inhibitors. Chem Biol 9: 3–16.
He L, He X, Lim LP, de Stanchina E, Xuan Z, Liang Y et al. (2007). A microRNA component of the p53 tumour suppressor network. Nature 447: 1130–1134.
Iorio MV, Visone R, Di Leva G, Donati V, Petrocca F, Casalini P et al. (2007). MicroRNA signatures in human ovarian cancer. Cancer Res 67: 8699–8707.
Izzotti A, Calin GA, Arrigo P, Steele VE, Croce CM, De Flora S . (2009). Downregulation of microRNA expression in the lungs of rats exposed to cigarette smoke. FASEB J 23: 806–812.
Koishi K, Yoshikawa R, Tsujimura T, Hashimoto-Tamaoki T, Kojima S, Yanagi H et al. (2006). Persistent CXCR4 expression after preoperative chemoradiotherapy predicts early recurrence and poor prognosis in esophageal cancer. World J Gastroenterol 12: 7585–7590.
Kozak M . (2008). Faulty old ideas about translational regulation paved the way for current confusion about how microRNAs function. Gene 423: 108–115.
Krishan A . (1975). Rapid flow cytofluorometric analysis of mammalian cell cycle by propidium iodide staining. J Cell Biol 66: 188–193.
Laemmli UK . (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227: 680–685.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al. (2005). MicroRNA expression profiles classify human cancers. Nature 435: 834–838.
Moskovits N, Kalinkovich A, Bar J, Lapidot T, Oren M . (2006). p53 attenuates cancer cell migration and invasion through repression of SDF-1/CXCL12 expression in stromal fibroblasts. Cancer Res 66: 10671–10676.
Nicolas FE, Pais H, Schwach F, Lindow M, Kauppinen S, Moulton V et al. (2008). Experimental identification of microRNA-140 targets by silencing and overexpressing miR-140. RNA 14: 2513–2520.
O'Brien CA, Pollett A, Gallinger S, Dick JE . (2007). A human colon cancer cell capable of initiating tumour growth in immunodeficient mice. Nature 445: 106–110.
Pillai RS, Bhattacharyya SN, Fillipowicz W . (2007). Repression of protein synthesis by miRNAs: how many mechanisms? Trends cell Biol 17: 118–126.
Plasterk RH . (2006). MicroRNAs in animal development. Cell 124: 877–881.
Raver-Shapira N, Meiri E, Spector Y, Rosenfeld N, Moskovits N, Bentwich Z et al. (2007). Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol Cell 26: 731–743.
Ricci-Vitiani L, Lombardi DG, Pilozzi E, Biffoni M, Todaro M, Peschle C et al. (2007). Identification and expansion of human colon-cancer-initiating cells. Nature 445: 111–115.
Sengupta N, Seto E . (2004). Regulation of histone deacetylase activities. J Cell Biochem 93: 57–67.
Song B, Wang Y, Kudo K, Gavin EJ, Xi Y, Ju J . (2008). miR-192 regulates dihydrofolate reductase and cellular proliferation through the p53–microRNA circuit. Clin Cancer Res 14: 8080–8086.
Tuddenham L, Wheeler G, Ntounia-Fousara S, Waters J, Hajihosseini MK, Clark I et al. (2006). The cartilage specific microRNA-140 targets histone deacetylase 4 in mouse cells. FEBS Lett 580: 4214–4217.
Volinia S, Calin GA, Liu CG, Ambs S, Cimmino A, Petrocca F et al. (2006). A microRNA expression signature of human solid tumors defines cancer gene targets. Proc Natl Acad Sci USA 103: 2257–2261.
Wilson AJ, Byun DS, Nasser S, Murray LB, Ayyanar K, Arango D et al. (2008). HDAC4 promotes growth of colon cancer cells via repression of p21. Mol Biol Cell 19: 4062–4075.
Yang XJ, Grégoire S . (2005). Class II histone deacetylases: from sequence to function, regulation, and clinical implication. Mol Cell Biol 25: 2873–2874.
Zhang B, Wang Q, Pan X . (2007). MicroRNAs and their regulatory roles in animals and plants. J Cell Physiol 210: 279–289.
Zou GM . (2008). Cancer initiating cells or cancer stem cells in the gastrointestinal tract and liver. J Cell Physiol 217: 598–604.
Acknowledgements
We appreciate the critical reading of the paper by Stephanie Burke (Stony Brook University). This work was supported by Stony Brook Translational Research Laboratory Start-up fund and NIH CA114043 (J Ju) and MH075020 (J Ju).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Oncogene website (http://www.nature.com/onc)
Supplementary information
Rights and permissions
About this article
Cite this article
Song, B., Wang, Y., Xi, Y. et al. Mechanism of chemoresistance mediated by miR-140 in human osteosarcoma and colon cancer cells. Oncogene 28, 4065–4074 (2009). https://doi.org/10.1038/onc.2009.274
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2009.274
Keywords
This article is cited by
-
miR-140-3P Induces Chemotherapy Resistance in Esophageal Carcinoma by Targeting the NFYA-MDR1 Axis
Applied Biochemistry and Biotechnology (2023)
-
MALAT1-miRNAs network regulate thymidylate synthase and affect 5FU-based chemotherapy
Molecular Medicine (2022)
-
c-Myb-mediated inhibition of miR-601 in facilitating malignance of osteosarcoma via augmentation of PKMYT1
Scientific Reports (2022)
-
MicroRNAs and drug resistance in colorectal cancer with special focus on 5-fluorouracil
Molecular Biology Reports (2022)
-
EVs delivery of miR-1915-3p improves the chemotherapeutic efficacy of oxaliplatin in colorectal cancer
Cancer Chemotherapy and Pharmacology (2021)