Key Points
-
DNA methylation is an epigenetic mark that is associated with gene silencing.
-
Novel high-throughput techniques have begun to generate much larger scale maps of DNA methylation than were previously available.
-
The new information has broadly confirmed several conclusions that were based on analysis of specific regions of the genome. For example, many promoters in mammalian genome are marked by unmethylated CpG islands.
-
This new data shows, however, that CpG islands are often found at intergenic regions and can acquire methylation in somatic cells. Evidence for tissue-specific CpG methylation patterns revives the idea that gene expression might be controlled by altered methylation of regulatory elements.
-
An unexpected finding is that the bodies of active genes in plants and animals are often heavily methylated. The function of gene-body methylation is presently unknown, but might serve to suppress intragenic transcription initiation.
-
Emerging high-throughput DNA sequencing technologies promise to provide complete DNA methylation maps that will transform our understanding of this epigenetic mark.
Abstract
The genomes of many animals, plants and fungi are tagged by methylation of DNA cytosine. To understand the biological significance of this epigenetic mark it is essential to know where in the genome it is located. New techniques are making it easier to map DNA methylation patterns on a large scale and the results have already provided surprises. In particular, the conventional view that DNA methylation functions predominantly to irreversibly silence transcription is being challenged. Not only is promoter methylation often highly dynamic during development, but many organisms also seem to target DNA methylation specifically to the bodies of active genes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Bird, A. Perceptions of epigenetics. Nature 447, 396–398 (2007).
Li, E., Bestor, T. H. & Jaenisch, R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 69, 915–926 (1992).
Okano, M., Bell, D. W., Haber, D. A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).
Chen, T. & Li, E. Structure and function of eukaryotic DNA methyltransferases. Curr. Top. Dev. Biol. 60, 55–89 (2004).
Karpf, A. R. & Matsui, S. Genetic disruption of cytosine DNA methyltransferase enzymes induces chromosomal instability in human cancer cells. Cancer Res. 65, 8635–8639 (2005).
Dodge, J. E. et al. Inactivation of Dnmt3b in mouse embryonic fibroblasts results in DNA hypomethylation, chromosomal instability, and spontaneous immortalization. J. Biol. Chem. 280, 17986–17991 (2005).
Walsh, C. P., Chaillet, J. R. & Bestor, T. H. Transcription of IAP endogenous retroviruses is constrained by cytosine methylation. Nature Genet. 20, 116–117 (1998).
Bird, A. P. & Wolffe, A. P. Methylation-induced repression — belts, braces, and chromatin. Cell 99, 451–454 (1999).
Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 16, 6–21 (2002).
Bird, A. P. CpG-rich islands and the function of DNA methylation. Nature 321, 209–213 (1986).
Selker, E. U. et al. The methylated component of the Neurospora crassa genome. Nature 422, 893–897 (2003).
Bird, A. P., Taggart, M. H. & Smith, B. A. Methylated and unmethylated DNA compartments in the sea urchin genome. Cell 17, 889–901 (1979).
Tweedie, S., Charlton, J., Clark, V. & Bird, A. Methylation of genomes and genes at the invertebrate–vertebrate boundary. Mol. Cell Biol. 17, 1469–1475 (1997).
Montero, L. M. et al. The distribution of 5-methylcytosine in the nuclear genome of plants. Nucleic Acids Res. 20, 3207–3210 (1992).
Palmer, L. E. et al. Maize genome sequencing by methylation filtration. Science 302, 2115–2117 (2003).
SanMiguel, P. et al. Nested retrotransposons in the intergenic regions of the maize genome. Science 274, 765–768 (1996).
Chan, S. W., Henderson, I. R. & Jacobsen, S. E. Gardening the genome: DNA methylation in Arabidopsis thaliana. Nature Rev. Genet. 6, 351–360 (2005).
Chan, S. W. et al. RNA silencing genes control de novo DNA methylation. Science 303, 1336 (2004).
Mette, M. F., Aufsatz, W., van der Winden, J., Matzke, M. A. & Matzke, A. J. Transcriptional silencing and promoter methylation triggered by double-stranded RNA. EMBO J. 19, 5194–5201 (2000).
Frommer, M. et al. A genomic sequencing protocol that yields a positive display of 5-methylcytosine residues in individual DNA strands. Proc. Natl Acad. Sci. USA 89, 1827–1831 (1992).
Eckhardt, F. et al. DNA methylation profiling of human chromosomes 6, 20 and 22. Nature Genet. 38, 1378–1385 (2006). In this paper, large-scale bisulphite sequence analysis of human chromosomal regions reveals greater variation in methylation patterns between tissues in a given individual than between individuals. Methylated regions were highly conserved, raising the possibility that they correspond to regulatory sequences.
Rollins, R. A. et al. Large-scale structure of genomic methylation patterns. Genome Res. 16, 157–163 (2006).
Bird, A. P. Use of restriction enzymes to study eukaryotic DNA methylation. II: the symmetry of methylated sites supports semi-conservative copying of the methylation pattern. J. Mol. Biol. 118, 48–60 (1978).
Schumacher, A. et al. Microarray-based DNA methylation profiling: technology and applications. Nucleic Acids Res. 34, 528–542 (2006).
Khulan, B. et al. Comparative isoschizomer profiling of cytosine methylation: the HELP assay. Genome Res. 16, 1046–1055 (2006).
Weber, M. et al. Chromosome-wide and promoter-specific analyses identify sites of differential DNA methylation in normal and transformed human cells. Nature Genet. 37, 853–862 (2005).
Cross, S. H., Charlton, J. A., Nan, X. & Bird, A. P. Purification of CpG islands using a methylated DNA binding column. Nature Genet. 6, 236–244 (1994).
Zhang, X. et al. Genome-wide high-resolution mapping and functional analysis of DNA methylation in Arabidopsis. Cell 126, 1189–1201 (2006). This reference and reference 35 are two of the first genome-wide analyses of DNA methylation, in which gene body-specific methylation is described.
Illingworth, R. et al. A novel CpG island set identifies tissue-specific methylation at developmental gene loci. PLoS Biol. 6, e22 (2008). This paper presents a large-scale analysis of human CpG islands that are purified using an unmethylated CpG-affinity column. Half of all CGIs are not coincident with annotated promoters and a significant fraction becomes methylated in normal somatic tissues, particularly at genes involved in development.
Fan, J. B. et al. Illumina universal bead arrays. Methods Enzymol. 410, 57–73 (2006).
Korshunova, Y. et al. Massively parallel bisulphite pyrosequencing reveals the molecular complexity of breast cancer-associated cytosine-methylation patterns obtained from tissue and serum DNA. Genome Res. 18, 19–29 (2008).
Taylor, K. H. et al. Ultradeep bisulfite sequencing analysis of DNA methylation patterns in multiple gene promoters by 454 sequencing. Cancer Res. 67, 8511–8518 (2007).
Zilberman, D. & Henikoff, S. Genome-wide analysis of DNA methylation patterns. Development 134, 3959–3965 (2007).
Cokus, S. J. et al. Shotgun bisulphite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452, 215–219 (2008).
Zilberman, D., Gehring, M., Tran, R. K., Ballinger, T. & Henikoff, S. Genome-wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nature Genet. 39, 61–69 (2007).
Henderson, I. R. & Jacobsen, S. E. Epigenetic inheritance in plants. Nature 447, 418–424 (2007).
Vaughn, M. W. et al. Epigenetic natural variation in Arabidopsis thaliana. PLoS Biol. 5, e174 (2007).
Simmen, M. W. et al. Non-methylated transposable elements and methylated genes in a chordate genome. Science 283, 1164–1167 (1999).
Suzuki, M. M., Kerr, A. R., De Sousa, D. & Bird, A. CpG methylation is targeted to transcription units in an invertebrate genome. Genome Res. 17, 625–631 (2007). In this paper, analysis of DNA methylation in the invertebrate chordate C. intestinalis establishes that about half of all genes are methylated and show gene-body methylation.
Field, L. M., Devonshire, A. L., Ffrench-Constant, R. H. & Forde, B. G. Changes in DNA methylation are associated with loss of insecticide resistance in the peach-potato aphid Myzus persicae (Sulz.). FEBS Lett. 243, 323–327 (1989).
Field, L. M. Methylation and expression of amplified esterase genes in the aphid Myzus persicae (Sulzer). Biochem. J. 349, 863–868 (2000).
Wang, Y. et al. Functional CpG methylation system in a social insect. Science 314, 645–647 (2006).
Rabinowicz, P. D. et al. Genes and transposons are differentially methylated in plants, but not in mammals. Genome Res. 13, 2658–2664 (2003).
Rakyan, V. K. et al. DNA methylation profiling of the human major histocompatibility complex: a pilot study for the human epigenome project. PLoS Biol. 2, e405 (2004).
Bird, A., Taggart, M., Frommer, M., Miller, O. J. & Macleod, D. A fraction of the mouse genome that is derived from islands of nonmethylated, CpG-rich DNA. Cell 40, 91–99 (1985).
Mohandas, T., Sparkes, R. S. & Shapiro, L. J. Reactivation of an inactive human X-chromosome: evidence for X-inactivation by DNA methylation. Science 211, 393–396 (1981).
Venolia, L. et al. Transformation with DNA from 5-azacytidine-reactivated X chromosomes. Proc. Natl. Acad. Sci. USA 79, 2352–2354 (1982).
Hellman, A. & Chess, A. Gene body-specific methylation on the active X chromosome. Science 315, 1141–1143 (2007). This study shows that genes on the active X chromosome are in fact more methylated than those on the inactive X chromosome, and that methylation is high within gene bodies.
Carrozza, M. J. et al. Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123, 581–592 (2005).
Vakoc, C. R., Mandat, S. A., Olenchock, B. A. & Blobel, G. A. Histone H3 lysine 9 methylation and HP1gamma are associated with transcription elongation through mammalian chromatin. Mol. Cell 19, 381–391 (2005).
Schaefer, C. B., Ooi, S. K., Bestor, T. H. & Bourc'his, D. Epigenetic decisions in mammalian germ cells. Science 316, 398–399 (2007).
Gehring, M. & Henikoff, S. DNA methylation dynamics in plant genomes. Biochim. Biophys. Acta 1769, 276–286 (2007).
Ma, Y. et al. DNA CpG hypomethylation induces heterochromatin reorganization involving the histone variant macroH2A. J. Cell Sci. 118, 1607–1616 (2005).
Slotkin, R. K. & Martienssen, R. Transposable elements and the epigenetic regulation of the genome. Nature Rev. Genet. 8, 272–285 (2007).
Wilson, A. S., Power, B. E. & Molloy, P. L. DNA hypomethylation and human diseases. Biochim. Biophys. Acta 1775, 138–162 (2007).
Bird, A. P. Gene number, noise reduction and biological complexity. Trends Genet. 11, 94–100 (1995).
Robertson, K. D. DNA methylation and human disease. Nature Rev. Genet. 6, 597–610 (2005).
Hsieh, C.-L. Evidence that protein binding specifies sites of DNA demethylation. Mol. Cell Biol. 19, 46–56 (1999).
Antequera, F. & Bird, A. Number of CpG islands and genes in human and mouse. Proc. Natl. Acad. Sci. USA 90, 11995–11999 (1993).
Cross, S., Kovarik, P., Schmidtke, J. & Bird, A. P. Non-methylated islands in fish genomes are GC-poor. Nucleic Acids Res. 19, 1469–1474 (1991).
Glass, J. L. et al. CG dinucleotide clustering is a species-specific property of the genome. Nucleic Acids Res. 35, 6798–6807 (2007).
Takai, D. & Jones, P. A. Comprehensive analysis of CpG islands in human chromosomes 21 and 22. Proc. Natl Acad. Sci. USA 99, 3740–3745 (2002).
Gardiner-Gardner, M. & Frommer, M. CpG islands in vertebrate genomes. J. Mol. Biol. 196, 261–282 (1987).
Panning, B. & Jaenisch, R. DNA hypomethylation can activate Xist expression and silence X-linked genes. Genes Dev. 10, 1991–2002 (1996).
Sleutels, F., Zwart, R. & Barlow, D. P. The non-coding Air RNA is required for silencing autosomal imprinted genes. Nature 415, 810–813 (2002).
Antequera, A. & Bird, A. in DNA Methylation: Molecular Biology and Biological Significance (eds J. P. Jost & H. P. Saluz) 169–185 (Birkhäuser, Basel, 1993).
Stoger, R. et al. Maternal-specific methylation of the imprinted mouse Igf2r locus identifies the expressed locus as carrying the imprinting signal. Cell 73, 61–71 (1993).
Sutcliffe, J. S. et al. Deletions of a differentially methylated CpG island at the SNRPN gene define a putative imprinting control region. Nature Genet. 8, 52–58 (1994).
Maatouk, D. M. et al. DNA methylation is a primary mechanism for silencing postmigratory primordial germ cell genes in both germ cell and somatic cell lineages. Development 133, 3411–3418 (2006).
Song, F. et al. Association of tissue-specific differentially methylated regions (TDMs) with differential gene expression. Proc. Natl Acad. Sci. USA 102, 3336–3341 (2005).
Yamada, Y. et al. A comprehensive analysis of allelic methylation status of CpG islands on human chromosome 21q. Genome Res. 14, 247–266 (2004).
Weber, M. et al. Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome. Nature Genet. 39, 457–466 (2007). This paper presents a large-scale analysis of DNA methylation at human promoters using microarrays. Variable methylation was particularly apparent at moderately CpG-rich promoters, suggesting a role in regulation of gene expression.
Petronis, A. Human morbid genetics revisited: relevance of epigenetics. Trends Genet. 17, 142–146 (2001).
Fraga, M. F. et al. Epigenetic differences arise during the lifetime of monozygotic twins. Proc. Natl Acad. Sci. USA 102, 10604–10609 (2005).
Ladd-Acosta, C. et al. DNA methylation signatures within the human brain. Am. J. Hum. Genet. 81, 1304–1315 (2007).
Siegmund, K. D. et al. DNA methylation in the human cerebral cortex is dynamically regulated throughout the life span and involves differentiated neurons. PLoS ONE 2, e895 (2007).
Jones, P. A. & Baylin, S. B. The epigenomics of cancer. Cell 128, 683–692 (2007).
Krieg, A. M. Therapeutic potential of Toll-like receptor 9 activation. Nature Rev. Drug Discov. 5, 471–484 (2006).
Hibino, T. et al. The immune gene repertoire encoded in the purple sea urchin genome. Dev. Biol. 300, 349–365 (2006).
Rountree, M. R. & Selker, E. U. DNA methylation inhibits elongation but not initiation of transcription in Neurospora crassa. Genes Dev. 11, 2383–2395 (1997).
Takata, M. et al. Rice transposable elements are characterized by various methylation environments in the genome. BMC Genomics 8, 469 (2007).
Bennetzen, J. L., Schrick, K., Springer, P. S., Brown, W. E. & SanMiguel, P. Active maize genes are unmodified and flanked by diverse classes of modified, highly repetitive DNA. Genome 37, 565–576 (1994).
Rabinowicz, P. D. et al. Differential methylation of genes and retrotransposons facilitates shotgun sequencing of the maize genome. Nature Genet. 23, 305–308 (1999).
Ashikawa, I. Gene-associated CpG islands in plants as revealed by analyses of genomic sequences. Plant J. 26, 617–625 (2001).
Goyon, C., Rossignol, J. L. & Faugeron, G. Native DNA repeats and methylation in Ascobolus. Nucleic Acids Res. 24, 3348–3356 (1996).
Galagan, J. E. & Selker, E. U. RIP: the evolutionary cost of genome defense. Trends Genet. 20, 417–423 (2004).
Selker, E. U. Premeiotic instability of repeated sequences in Neurospora crassa. Annu. Rev. Genet. 24, 579–613 (1990).
Faugeron, G. Diversity of homology-dependent gene silencing strategies in fungi. Curr. Opin. Microbiol. 3, 144–148 (2000).
Rhounim, L., Rossignol, J. L. & Faugeron, G. Epimutation of repeated genes in Ascobolus immersus. EMBO J. 11, 4451–4457 (1992).
Salzberg, A., Fisher, O., Siman-Tov, R. & Ankri, S. Identification of methylated sequences in genomic DNA of adult Drosophila melanogaster. Biochem. Biophys. Res. Commun. 322, 465–469 (2004).
Macleod, D., Clark, V. H. & Bird, A. Absence of genome-wide changes in DNA methylation during development of the zebrafish. Nature Genet. 23, 139–140 (1999).
Stancheva, I., El-Maarri, O., Walter, J., Niveleau, A. & Meehan, R. R. DNA methylation at promoter regions regulates the timing of gene activation in Xenopus laevis embryos. Dev. Biol. 243, 155–165 (2002).
Estecio, M. R. et al. LINE-1 hypomethylation in cancer is highly variable and inversely correlated with microsatellite instability. PLoS ONE 2, e399 (2007).
Yoder, J. A., Walsh, C. P. & Bestor, T. H. Cytosine methylation and the ecology of intragenomic parasites. Trends Genet. 13, 335–340 (1997).
Hatada, I., Hayashizaki, Y., Hirotsune, S., Komatsubara, H. & Mukai, T. A genomic scanning method for higher organisms using restriction sites as landmarks. Proc. Natl Acad. Sci. USA 88, 9523–9527 (1991).
Irizarry, R. A. et al. Comprehensive high-throughput arrays for relative methylation (CHARM). Genome Res. (2008).
Lippman, Z., Gendrel, A. V., Colot, V. & Martienssen, R. Profiling DNA methylation patterns using genomic tiling microarrays. Nature Methods 2, 219–224 (2005).
Keshet, I. et al. Evidence for an instructive mechanism of de novo methylation in cancer cells. Nature Genet. 38, 149–153 (2006).
Jorgensen, H. F., Adie, K., Chaubert, P. & Bird, A. P. Engineering a high-affinity methyl-CpG-binding protein. Nucleic Acids Res. 34, e96 (2006).
Dalma-Weiszhausz, D. D., Warrington, J., Tanimoto, E. Y. & Miyada, C. G. The affymetrix GeneChip platform: an overview. Methods Enzymol. 410, 3–28 (2006).
Nuwaysir, E. F. et al. Gene expression analysis using oligonucleotide arrays produced by maskless photolithography. Genome Res. 12, 1749–1755 (2002).
Reinders, J. et al. Genome-wide, high-resolution DNA methylation profiling using bisulfite-mediated cytosine conversion. Genome Res. 18, 469–476 (2008).
Yuan, E. et al. A single nucleotide polymorphism chip-based method for combined genetic and epigenetic profiling: validation in decitabine therapy and tumor/normal comparisons. Cancer Res. 66, 3443–3451 (2006).
Bentley, D. R. Whole-genome re-sequencing. Curr. Opin. Genet. Dev. 16, 545–552 (2006).
Rauch, T. A. et al. High-resolution mapping of DNA hypermethylation and hypomethylation in lung cancer. Proc. Natl Acad. Sci. USA 105, 252–257 (2008).
Bibikova, M. et al. High-throughput DNA methylation profiling using universal bead arrays. Genome Res. 16, 383–393 (2006).
Acknowledgements
This work was supported by a grant from the Wellcome Trust. We thank A. Deaton for constructive criticism.
Author information
Authors and Affiliations
Corresponding author
Related links
Glossary
- Imprinted gene
-
A gene that is expressed or silenced depending on which parent contributed it to the zygote. In a mouse cell, for example, the paternal insulin-like growth factor allele is expressed, but the maternal allele is not. In some cases, imprinting depends on differential DNA methylation of gene regulatory regions.
- CpG island
-
(CGI). A DNA patch of approximately 1,000 bp, within which the dinucleotide CpG occurs at close to its expected frequency. This contrasts with the majority of the vertebrate genome, in which CpG is depleted. Despite the abundance of CpGs that could potentially be methylated, CGIs are unmethylated in germ cells and most are also DNA methylation free in somatic cells. In mammals, CGIs are GC-rich in base composition (∼65%) compared with the genome as a whole (∼40%).
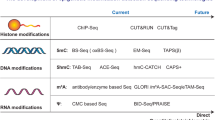
- 454 sequencing and Solexa bisulphite sequencing
-
Independent proprietary high-throughput DNA-sequencing technologies that both use massively parallel sequencing-by-synthesis approaches. These new methods allow an increase in generated sequence per run of about two orders of magnitude compared with conventional Sanger sequencing technologies, and therefore allow rapid comprehensive sequence screening of large genomic fractions or whole genomes.
- Heterochromatic knob
-
A chromosomal region that can be identified microscopically as being darkly stained compared with surrounding chromatin. DNA sequence analysis has shown that knobs often contain highly repeated DNA sequences. They were described initially in the 1930's by McClintock during her studies of maize chromosome structure.
Rights and permissions
About this article
Cite this article
Suzuki, M., Bird, A. DNA methylation landscapes: provocative insights from epigenomics. Nat Rev Genet 9, 465–476 (2008). https://doi.org/10.1038/nrg2341
Issue Date:
DOI: https://doi.org/10.1038/nrg2341
This article is cited by
-
Discovery of novel DNA methylation biomarker panels for the diagnosis and differentiation between common adenocarcinomas and their liver metastases
Scientific Reports (2024)
-
DNMT3B PWWP mutations cause hypermethylation of heterochromatin
EMBO Reports (2024)
-
High temperature influences DNA methylation and transcriptional profiles in sea urchins (Strongylocentrotus intermedius)
BMC Genomics (2023)
-
Mediation effects of DNA methylation and hydroxymethylation on birth outcomes after prenatal per- and polyfluoroalkyl substances (PFAS) exposure in the Michigan mother–infant Pairs cohort
Clinical Epigenetics (2023)
-
Mastering DNA chromatogram analysis in Sanger sequencing for reliable clinical analysis
Journal of Genetic Engineering and Biotechnology (2023)