Abstract
Chromatographic retention factor log kIAM obtained from immobilized artificial membrane (IAM) HPLC with buffered, aqueous mobile phases and calculated molecular descriptors (molecular weight — log MW; molar volume — VM; polar surface area — PSA; total count of nitrogen and oxygen atoms -(N + O); count of freely rotable bonds — FRB; H-bond donor count — HD; H-bond acceptor count — HA; energy of the highest occupied molecular orbital — EHOMO; energy of the lowest unoccupied orbital — ELUMO; dipole moment — DM; polarizability — α) obtained for a group of 175 structurally unrelated compounds were tested in order to generate useful models of solutes’ soil-water partition coefficient normalized to organic carbon log Koc. It was established that log kIAM obtained in the conditions described in this study is not sufficient as a sole predictor of the soil-water partition coefficient. Simple, potentially useful models based on log kIAM and a selection of readily available, calculated descriptors and accounting for over 88% of total variability were generated using multiple linear regression (MLR) and artificial neural networks (ANN). The models proposed in the study were tested on a group of 50 compounds with known experimental log Koc values by plotting the calculated vs. experimental values. There is a good close similarity between the calculated and experimental data for both MLR and ANN models for compounds from different chemical families (R2 ≥ 0.80, n = 50) which proves the models’ reliability.
Similar content being viewed by others
Introduction
The fate and transport of solutes in the environment depend on their physico-chemical properties such as lipophilicity, volatility, water solubility, and ability to partition between soil and water. The soil-water partition coefficient Koc (normalized to the soil organic carbon content to reduce the differences among soils) is a very important parameter governing the fate of compounds in the soil-water compartment (Doucette 2003). It influences the chemicals’ mobility in soil, their environmental persistence, and the processes of their removal in wastewater management facilities (Andrić et al. 2016). A very high value of Koc means a compound is strongly retained by soil and organic matter and does not move throughout the soil. A very low value means a molecule is highly mobile in soil. Koc is a very important input parameter for estimating environmental distribution and environmental risk related to a chemical substance. A highly mobile substance is more likely to move to groundwater or surface water from soil.
Direct determination of Koc is based on studies of partitioning phenomena in biphasic soil-water systems using batch or continuous flow experiments. Such tests are, however, tedious and time-consuming, and the results tend to be inconsistent due to experimental difficulties — incomplete separation of phases and volatilization and biological or chemical degradation of solutes (Hodson and Williams 1988). The need to obtain Koc values very rapidly and for large groups of compounds has led to the development of other methods. Entirely computational approaches are based on molecular descriptors such as water solubility WS (Gawlik et al. 1997), octanol-water partition coefficient Pow (Karickhoff et al. 1979; Hodson and Williams 1988; Pussemier et al. 1990; Müller and Kördel 1996; Gawlik et al. 1997; Doucette 2003), molecular connectivity indices MCI (Gawlik et al. 1997; Tao et al. 2001), topological indices (Meylan et al. 1992; Gawlik et al. 1997), or solvation parameters (Gawlik et al. 1997; Poole and Poole 1999; Nguyen et al. 2005; Poole et al. 2013). Alternatively, it is possible to predict soil-water partition using chromatographic descriptors obtained by liquid chromatography on octadecyl-, cyano-, diol-, ethyl-, trimethylammonium-, or aminopropyl-modified silica or on sorbents containing immobilized humic acid or soil (Helling and Turner 1968; Praveen-Kumar et al. 1987; Vowles and Mantoura 1987; Jamet and Thoisy-Dur 1988; Pussemier et al. 1990; Szabo et al. 1990a, b; Kördel et al. 1993, 1995a, b, 1997; Christianson and Howard 1994; Müller and Kördel 1996; Gawlik et al. 1997, 1998, 2000; Poole and Poole 1999; Szabó et al. 1999; Xu et al. 1999, 2002; Ravanel et al. 1999; Guo et al. 2004; Bermúdez-Saldaña et al. 2006; Mrozik et al. 2008; Andrić et al. 2010; Bi et al. 2010; Hidalgo-Rodríguez et al. 2012; Poole et al. 2013; Sobańska 2021). Chromatographic methods of Koc determination are fast and relatively cheap, with the benefit of high reproducibility, especially when commercially available stationary phases are used. It is generally accepted that log Koc is connected with the chromatographic retention factor log k via a Collander-type linear relationship: log Koc = a + b log k (Bermúdez-Saldaña et al. 2006). Linear relationships have also been discovered between log Koc and thin-layer chromatographic descriptors: RM value defined by Bate-Smith and Westall: RM = log (1/Rf − 1) (Bate-Smith and Westall 1950) and measured for a single composition of a mobile phase; RM0 value obtained from a series of chromatographic experiments with mobile phases containing different concentrations φ of a water-miscible solvent (organic modifier) by extrapolation of RM vs. φ plots to zero concentration of the modifier (most frequently by using the linear Soczewiński-Wachmeister equation: RM = RM0 + S φ (Soczewiński and Wachtmeister 1962)); the slope S taken from the Soczewiński-Wachmeister equation; the real or hypothetical concentration of the organic modifier furnishing RM = 0 (i.e., Rf = 0.5): φ0 = −RM0/S; PC1 (the 1st principal component computed for thin layer chromatographic retention data). Chromatographic retention parameters are usually considered to be sufficiently good as sole predictors of log Koc, but some authors suggest that other descriptors (e.g., polar surface area, PSA) should be incorporated to obtain better fitting models (Sobańska 2021).
Biomimetic liquid chromatography on immobilized artificial membrane (IAM) stationary phases has been used to predict the physicochemical properties and bioavailability of compounds for several years (Sobanska and Brzezinska 2017). IAM chromatographic phases are excellent tools to model drug distribution because of their ability to mimic natural membranes, and the main applications of IAM chromatography are in drug discovery (Valko 2019). The more recent applications of IAM chromatography extend from medicinal to environmental chemistry. Studies of compounds’ bioconcentration in aquatic organisms (Tsopelas et al. 2017, 2018), ecotoxicity of pesticides (Stergiopoulos et al. 2019), and soil-water sorption of herbicides (Hidalgo-Rodríguez et al. 2012) prove that IAM chromatography is a source of valuable information in many areas of research. The objective of the present study was to propose useful models of the soil-water partition coefficients normalized to organic carbon (Koc) using a large group of structurally diverse organic compounds, based their IAM chromatographic and computational descriptors. The Koc models considered in this study should be applicable during early steps of a drug discovery process, when drugs’ physico-chemical and biological properties are often studied in vitro using IAM chromatography. At this stage, high throughput is more important than accuracy and the chromatographic and computational data are collected for other purposes. It was anticipated that IAM chromatographic and computational studies of soil-water partition of compounds could be carried out concurrently with pharmacokinetic studies.
Experimental
Calculated molecular descriptors
The molecular descriptors for compounds investigated during this study were calculated with HyperChem 8.0, utilizing PM3 semi-empirical method with Polak-Ribiere conjugate gradient algorithm for semi-empirical structure optimization (Polak and Ribiere 1969): total dipole moment — DM (D), molecular weight — Mw (g mol−1), energy of the highest occupied molecular orbital — EHOMO (eV), and energy of the lowest unoccupied molecular orbital − ELUMO (eV). Other physicochemical parameters (octanol-water partition coefficient — log Kow, polar surface area — PSA (Å2), H-bond donor count — HD, H-bond acceptor count — HA, polarizability — α (Å3), molar volume — VM (cm3), freely rotable bond count — FRB) were calculated using ACD/Labs 8.0 software, using the SMILES codes of molecules as the input data. (N + O) — total nitrogen and oxygen atom count was calculated from the molecular structures. The Abraham’s solvation parameters (A — hydrogen bond acidity; B — hydrogen bond basicity; S — dipolar interactions; E — excess molar refractivity; V — McGowan’s molecular volume) were taken from the paper by Sprunger et al. (2007). The calculated molecular descriptors are given in Table 1 (Supplementary Materials).
IAM chromatography
The chromatographic retention factors log kIAM used in this study were compiled by Sprunger at al. (Sprunger et al. 2007). They were obtained on a IAM.PC.DD2 HPLC column using an aqueous mobile phase buffered at pH ≤ 3 for carboxylic acids (to suppress their ionization) and in the pH range 6.5 to 7.5 for other compounds.
MLR and ANN models
Multiple linear regression (MLR) models were generated using Statistica v. 13, stepwise forward regression mode (F to enter set at 1 and F to remove set at 0).
Multilayer perceptron (MLP) networks, with the number of inputs the same as the number of variables, the varying number of hidden units, and one output unit, were generated using Statistica v. 13 (regression mode, Automated Network Search — ANS module, 1000 networks to train, 5 networks to retain). The neuron activation functions were selected from the following group: identity, logistic, hyperbolic tangent, and exponential. BFGS (Broyden-Fletcher-Goldfarb-Shanno algorithm) was used to train the network together with the sum of squares (SOS) error function.
Results and discussion
Calculated log Koc coefficients
A soil-water partition coefficient normalized to the organic carbon content (Koc) is an important parameter influencing the fate of solutes in the soil-water compartment. In this study, because of the lack of experimental data for the whole studied group of compounds, Koc was initially calculated using the four most commonly accepted computational approaches (Andrić et al. 2016) (Table 2, Supplementary Materials):
where MCI is the first-order molecular connectivity index; log Kow is the estimated octanol-water partition coefficient (KOWWIN); and A, B, S, E, and V are the Abraham’s solvation parameters (A — hydrogen bond acidity; B — hydrogen bond basicity; S — dipolar interactions; E — excess molar refractivity; V — McGowan’s molecular volume). Log Koc(1) and log Koc(2) values were calculated using the EpiSuite software (EpiWeb 4.1, KOCWIN module) (US EPA O 2015).
At this point, attention turned to log Koc(4) as a reference soil-water partition coefficient value readily available for the whole group of studied solutes. The known theoretical log Koc models (1) to (4) have their limitations: e.g., linear models based on log Kow (log Koc(2)) are not particularly useful for polar or ionizable solutes (Franco and Trapp 2008). However, Eq. (4) appears to be applicable to compounds across a relatively wide range of chemical families, including ionizable or polar molecules (e.g., ionizable or polar compounds within the studied group, whose experimental log Koc values are available (US EPA O 2015) — ethanol, 35; acetic acid, 98; phenylacetic acid, 122 and benzoic acid, 144).
Calculated log Koc coefficients vs. IAM chromatographic descriptors
The values of log Koc calculated according to Eq. (4) were plotted against the IAM retention factor log kIAM. The linear relationship between log Koc(4) and log kIAM (Eq. (5a)) accounts for 81% of total log Koc variability.
The results of log Koc modeling using a single chromatographic descriptor (log kIAM) obtained in this study (Eq. (5a)) are slightly poorer than those reported by Hidalgo-Rodríguez et al. (2012), who found the IAM retention factor superior to other chromatographic measures of soil-water partition. However, Hidalgo-Rodrigues et al. generated their IAM-chromatographic model (Eq. (5b)) using a significantly smaller group of compounds (n = 38) of much more limited structural diversity:
The group of compounds considered in the current study contains solutes of very different physico-chemical properties, e.g., molecules that are relatively strong organic acids and bases, from different chemical families and of different usage. The studied group contains simple organic chemicals (e.g., solvents), drugs (including, e.g., steroids, non-steroid anti-inflammatory drugs, topical anesthetics, beta-blockers, antibacterial and antiparasitic drugs), and components of fragrances/essential oils. Some of the studied compounds are likely to be released to the environment (e.g., by washing off the skin or with feces) and seriously affect the wildlife (e.g., by interacting with the reproductive processes of aquatic animals).
Multiple linear regression studies
In the current study, compared to the previous investigations (Hidalgo-Rodríguez et al. 2012), the focus is on the possibility of making the chromatographic method of Koc prediction using the IAM retention factor more universal by incorporating additional, easily calculated molecular descriptors, and by considering both linear and non-linear relationships between the independent variables and the studied property, rather than on selecting the most effective chromatographic system.
Equation (5a) accounts for only 81% of total log Koc variability; attempts were therefore made to improve a model by incorporating additional independent variables. However, this was not an easy task, since log kIAM already encodes some properties, responsible for partition and transport phenomena (especially lipophilicity expressed as log Kow — the correlation between log Kow and log kIAM is linear, with R2 = 0.84). When lipophilicity and redundant variables defining the molecular size were excluded from the analysis, Eq. (6) containing the following independent variables: log kIAM, log Mw, (N + O), ELUMO, α, EHOMO, HA, PSA, HD, and DM was generated by forward stepwise multiple regression (Fig. 1):
The group of 175 studied compounds was divided into two subsets: a training set (n = 125) and a test set (n = 50, compounds with known log Kocexp values). Equation (7), generated for the training set, and containing the same variables as Eq. (6), is as follows:
The values of log Koc(7) were calculated for the test set according to Eq. (7) and plotted against the reference log Koc(4) values to furnish a linear relationship (R2 = 0.86, or R2 = 0.93 when thiourea was removed as an outlier). The model (6) was also tested on the same test set of 50 compounds, whose log Kocexp values were available (US EPA O 2015). The resulting relationship between log Koc(6) and log Kocexp is linear, with R2 = 0.85.
Equation (6), despite encouraging results of validation, was found unsatisfying because it seems unpractical and over-parameterized — it contains 10 independent variables whose contributions (apart from log kIAM and log Mw) are not very significant. At this point, it was decided to simplify Eq. (6) as much as possible; the attempts to achieve it furnished Eqs. (8), (9), and (10), based on reduced sets of independent variables (Figs. 2, 3 and 4).
The group of 175 studied compounds was divided into two subsets: a training set (n = 125) and a test set (n = 50, compounds with known log Kocexp values). Equations (11), (12), and (13) generated for the training set, and containing the same variables as Eqs. (8), (9), and (10) are as follows:
The relationships between the log Koc values calculated for the test set according to Eqs. (11), (12), and (13) and the reference values — log Koc(4) are linear (R2 = 0.90, 0.89, and 0.90, respectively). The models (8), (9), and (10) were also tested on the same test set of 50 compounds, whose log Kocexp values were available (US EPA O 2015). The resulting relationships between the predicted and experimental log Koc values are linear, with R2 = 0.83, 0.80, and 0.80, respectively.
Equation (9) contains, apart from log kIAM and log MW, the total N and O atom count (N + O). This descriptor is a relatively good measure of polarity and H-bonding activity of molecules, and it has been used to predict their ADME properties, e.g., the blood-brain barrier permeability (Clark 2003) or skin permeability (Sobańska et al. 2021). However, it plays a rather minor role in soil-water partition modeling using MLR and Eq. (10) seems to be sufficient for rapid log Koc predictions.
Artificial neural network analysis
At this point, attention turned to artificial neural network (ANN) models. Artificial neural networks are widely used to predict drugs’ bioavailability (Carracedo-Reboredo et al. 2021) or properties such as an affinity for phospholipids using IAM chromatography and calculated descriptors (Ciura et al. 2021). The great advantages of neural networks compared to MLR are the possibility of utilizing both linear and non-linear relationships between input data and a predicted parameter and the ability of ANNs to learn these relationships directly from the data being modeled.
In this study, the ANN models were built for the same group of compounds (n = 175) that were used in the MLR analysis. 125 substances without the known log Kocexp data were used as a training set; the remaining compounds, whose log Kocexp were available (US EPA O 2015) were divided into a test set (n = 25) and a validation set (n= 25).
ANNs make it possible to process a large number of descriptors that can be easily obtained using readily available software. The selection of ANN input data is an important step because if the number of parameters is excessive considering the number of cases, overfitting may occur. An over-fitted model fits perfectly the data used as a training set but is likely to be inapplicable to additional cases (a simple example of an over-fitted model is a polynomial of a degree equal or close to the number of data points which fits the existing data very well but is not very good at generalization while extrapolated beyond the fitted data). In this study, a decision was made to consider, apart from the IAM retention factor log kIAM, a relatively small number of parameters, especially those that are known to be good predictors of environmental phenomena (Mamy et al. 2015). According to Mammy et al., 686 different molecular descriptors are found in 790 QSAR equations generated to predict 90 environmental parameters, but the descriptors that are most widely used are EHOMO, ELUMO, Mw, α, and DM. The input data used in this part of the study were selected from the following group, involved earlier in multiple regression analysis: log kIAM, log Mw, PSA, (N + O), HD, HA, α, DM, EHOMO, and ELUMO, and the predicted variable was log Koc(4). In the case of every ANN model, 1000 networks were generated and five with the smallest error were retained for further examination in search for a network that would give the results being in the best agreement with the reference data (log Koc(4)) for the whole group of studied compounds (n = 175) and with the experimental data (log Kocexp) for a subgroup of cases (n = 50). The ANN models investigated at this stage of the study involved the following sets of independent variables, used earlier in the multiple regression analysis (Eqs. (6), (8), (9), and (10)) (Table 4, Supplementary Materials):
-
ANN1: log kIAM, log Mw, PSA, (N + O), HD, HA, α, DM, EHOMO, ELUMO
-
ANN2: log kIAM, log Mw, (N + O), α, EHOMO, ELUMO
-
ANN3: log kIAM, log Mw, (N + O)
-
ANN4: log kIAM, log Mw
The details of all the ANNs considered at this stage of the study are given in Table 4 (Supplementary Materials). The networks ANN1 to ANN4 were studied using a tool known as global sensitivity analysis (GSA), which rates the importance of the models’ input variable by computing sums of squared residuals for the model when the respective predictor is eliminated, compared to the full model. When an input variable scores 1 or less in GSA, it means that this particular network would perform better if this variable is omitted; however, no such situation occurred for the networks ANN1 to ANN4. All the 20 most promising networks were analyzed by plotting the predicted log KocANN values against the reference log Koc(4) values obtained for the whole group of 175 studied compounds. The predicted log KocANN values were also plotted against the experimental log Kocexp values for 50 compounds, whose log Kocexp values are available. The resulting dependences between log KocANN and log Koc(4) and between log KocANN and log Kocexp are linear (see Table 4, Supplementary Materials). It was found that the networks of the ANN2 family have the predictive abilities comparable to those of ANN1 networks, but since they involve a smaller number of input variables and hidden layers, they are likely to be less prone to overfitting. ANN4 networks, based on just two input variables (log kIAM and log Mw), give the prediction results comparable to those obtained using the ANN3 networks; similar to MLR models (Eqs. (9) and (10)), it seems that just two variables, log kIAM and log Mw, account for ca. 88–89% of the total log Koc variability (depending on the ANN’s architecture).
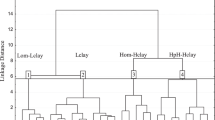
The differences and similarities between the reference values (log Koc(4)) and the predicted values obtained according to MLR and ANN models described above were studied briefly using Cluster Analysis (k-means method). 25 log Koc values (log Koc(4) and the values predicted using MLR and ANN models: log Koc(6), log Koc(8) to log Koc(10), log KocANN1, log KocANN2, log KocANN3, and log KocANN4) obtained for 175 compounds were separated between 5 clusters with minimum within-cluster variances and maximum variances between clusters (Table 5, Supplementary Materials). It was established that the reference values (log Koc(4)) form a cluster with the values calculated using ANN2-1, ANN2-2, ANN2-3, and ANN2-4 networks (Cluster 5). The mean log Koc values obtained for clusters 1 to 5 were plotted against the experimental log Kocexp values for 50 compounds whose experimental log Kocexp values are available. The resulting relationships are linear (R2 = 0.84, 0.79, 0.85, 0.80, and 0.84, respectively). The differences between the prediction results for clusters 1 to 5 are relatively small (Fig. 5); for the great majority of studied compounds 1 to 175, the mean values of log Koc calculated for all the clusters are similar (S.D. below 0.15, Table 6, Supplementary Materials). However, a network that attracts particular attention is ANN2-2 (Fig. 6) based on 6 input variables and giving the prediction results superior to those obtained using the MLR model involving the same set of independent variables (Eq. (8)). The values of log KocANN2-2 calculated using this network are in agreement with the reference values log Koc(4) (R2 = 0.95, n = 175) and with the experimental values log Kocexp (R2 = 0.86, n = 50).
Conclusions
In this study, new models of the soil-water partition coefficient Koc were developed using IAM chromatographic retention factors and calculated physico-chemical parameters of 175 structurally diverse compounds. The relationships between log Koc and the independent variables were either linear (multiple linear regression, MLR models) or non-linear (artificial neural network, ANN models).
Due to the limited availability of experimental permeability data for the solutes investigated in this study, the reference soil-water partition coefficients were calculated according to a widely accepted theoretical model based on Abraham’s solvation parameters. The values of log Koc obtained using this model are in agreement with the experimental data (where available).
Both MLR and ANN models proposed in this study have certain advantages. Compared to the MLR model, the ANN model gives slightly better prediction results which indicate non-linearity and complexity of correlations between the studied descriptors and the soil-water partition coefficient. The main descriptors, responsible for the variability of log Koc in MLR equations, are log kIAM and log Mw. When other independent variables are incorporated, the ANN models gain more in predictive power than MLR equations. Non-linear modeling is promising in terms of predicting compounds’ mobility in soil with high accuracy. However, linear relationships (Eq. (10)) give sufficiently good results and have the benefit of simplicity. Linear models are also more intuitive and can be easily applied during early steps of a drug discovery process. At this stage, high throughput is essential; IAM chromatographic and computational descriptors are collected to study drugs’ pharmacokinetic properties, so acquisition of input data for soil mobility studies would not require any additional effort. IAM chromatographic and computational studies of soil-water partition of compounds may therefore be a valuable extension of ADMET studies.
Data availability
Data used in this manuscript are given as Supplementary Materials or in references cited in the text.
References
Andrić FLJ, Trifković JĐ, Radoičić AD et al (2010) Determination of the soil–water partition coefficients (logKOC) of some mono- and poly-substituted phenols by reversed-phase thin-layer chromatography. Chemosphere 81:299–305. https://doi.org/10.1016/j.chemosphere.2010.07.049
Andrić F, Šegan S, Tešić Ž, Milojković-Opsenica D (2016) Chromatographic methods in determination of the soil–water partition coefficient. J Liq Chromatogr Relat Technol 39:249–256. https://doi.org/10.1080/10826076.2016.1163173
Bate-Smith EC, Westall RG (1950) Chromatographic behaviour and chemical structure I. Some naturally occuring phenolic substances. Biochim Biophys Acta 4:427–440. https://doi.org/10.1016/0006-3002(50)90049-7
Bermúdez-Saldaña JM, Escuder-Gilabert L, Medina-Hernández MJ et al (2006) Chromatographic estimation of the soil-sorption coefficients of organic compounds. TrAC Trends Anal Chem 25:122–132. https://doi.org/10.1016/j.trac.2005.09.004
Bi E, Schmidt TC, Haderlein SB (2010) Practical issues relating to soil column chromatography for sorption parameter determination. Chemosphere 80:787–793. https://doi.org/10.1016/j.chemosphere.2010.05.006
Carracedo-Reboredo P, Liñares-Blanco J, Rodríguez-Fernández N et al (2021) A review on machine learning approaches and trends in drug discovery. Comput Struct Biotechnol J 19:4538–4558. https://doi.org/10.1016/j.csbj.2021.08.011
Christianson CB, Howard RG (1994) Use of soil thin-layer chromatography to assess the mobility of the phosphoric triamide urease inhibitors and urea in soil. Soil Biol Biochem 26:1161–1164. https://doi.org/10.1016/0038-0717(94)90138-4
Ciura K, Kovačević S, Pastewska M et al (2021) Prediction of the chromatographic hydrophobicity index with immobilized artificial membrane chromatography using simple molecular descriptors and artificial neural networks. J Chromatogr A 1660:462666. https://doi.org/10.1016/j.chroma.2021.462666
Clark DE (2003) In silico prediction of blood–brain barrier permeation. Drug Discov Today 8:927–933. https://doi.org/10.1016/S1359-6446(03)02827-7
Doucette WJ (2003) Quantitative structure-activity relationships for predicting soil-sediment sorption coefficients for organic chemicals. Environ Toxicol Chem 22:1771–1788. https://doi.org/10.1897/01-362
Franco A, Trapp S (2008) Estimation of the soil–water partition coefficient normalized to organic carbon for ionizable organic chemicals. Environ Toxicol Chem 27:1995–2004. https://doi.org/10.1897/07-583.1
Gawlik BM, Sotiriou N, Feicht EA et al (1997) Alternatives for the determination of the soil adsorption coefficient, KOC, of non-ionicorganic compounds — a review. Chemosphere 34:2525–2551. https://doi.org/10.1016/S0045-6535(97)00098-2
Gawlik BM, Feicht EA, Karcher W et al (1998) Application of the European reference soil set (EUROSOILS) to a HPLC-screening method for the estimation of soil adsorption coefficients of organic compounds. Chemosphere 36:2903–2919. https://doi.org/10.1016/S0045-6535(97)10247-8
Gawlik BM, Kettrup A, Muntau H (2000) Estimation of soil adsorption coefficients of organic compounds by HPLC screening using the second generation of the European reference soil set. Chemosphere 41:1337–1347. https://doi.org/10.1016/S0045-6535(00)00015-1
Guo R, Liang X, Chen J et al (2004) Prediction of soil organic carbon partition coefficients by soil column liquid chromatography. J Chromatogr A 1035:31–36. https://doi.org/10.1016/j.chroma.2004.02.043
Helling CS, Turner BC (1968) Pesticide mobility: determination by soil thin-layer chromatography. Science 162:562–563. https://doi.org/10.1126/science.162.3853.562
Hidalgo-Rodríguez M, Fuguet E, Ràfols C, Rosés M (2012) Performance of chromatographic systems to model soil–water sorption. J Chromatogr A 1252:136–145. https://doi.org/10.1016/j.chroma.2012.06.058
Hodson J, Williams NA (1988) The estimation of the adsorption coefficient (Koc) for soils by high performance liquid chromatography. Chemosphere 17:67–77. https://doi.org/10.1016/0045-6535(88)90045-8
Jamet P, Thoisy-Dur J-C (1988) Pesticide mobility in soils: assessment of the movement of isoxaben by soil thin-layer chromatography. Bull Environ Contam Toxicol 41:135–142. https://doi.org/10.1007/BF01689070
Karickhoff SW, Brown DS, Scott TA (1979) Sorption of hydrophobic pollutants on natural sediments. Water Res 13:241–248. https://doi.org/10.1016/0043-1354(79)90201-X
Kördel W, Stutte J, Kotthoff G (1993) HPLC-screening method for the determination of the adsorption-coefficient on soil-comparison of different stationary phases. Chemosphere 27:2341–2352. https://doi.org/10.1016/0045-6535(93)90257-6
Kördel W, Kotthoff G, Müller J (1995a) HPLC-screening method for the determination of the adsorption-coefficient on soil — results of a ring-test. Chemosphere 30:1373–1384. https://doi.org/10.1016/0045-6535(95)00030-C
Kördel W, Stutte J, Kotthoff G (1995b) HPLC-screening method to determine the adsorption coefficient in soil-comparison of immobilized humic acid and clay mineral phases for cyanopropyl columns. Sci Total Environ 162:119–125. https://doi.org/10.1016/0048-9697(95)04443-5
Kördel W, Hennecke D, Herrmann M (1997) Application of the HPLC-screening method for the determination of the adsorption coefficient on sewage sludges. Chemosphere 35:121–128. https://doi.org/10.1016/S0045-6535(97)00144-6
Mamy L, Patureau D, Barriuso E et al (2015) Prediction of the fate of organic compounds in the environment from their molecular properties: a review. Crit Rev Environ Sci Technol 45:1277–1377. https://doi.org/10.1080/10643389.2014.955627
Meylan W, Howard PH, Boethling RS (1992) Molecular topology/fragment contribution method for predicting soil sorption coefficients. Environ Sci Technol 26:1560–1567. https://doi.org/10.1021/es00032a011
Mrozik W, Nichthauser J, Stepnowski P (2008) Prediction of the adsorption coefficients for imidazolium ionic liquids in soils using cyanopropyl stationary phase. Pol J Environ Stud 17:383–388
Müller M, Kördel W (1996) Comparison of screening methods for the estimation of adsorption coefficients on soil. Chemosphere 32:2493–2504. https://doi.org/10.1016/0045-6535(96)00148-8
Nguyen TH, Goss K-U, Ball WP (2005) Polyparameter linear free energy relationships for estimating the equilibrium partition of organic compounds between water and the natural organic matter in soils and sediments. Environ Sci Technol 39:913–924. https://doi.org/10.1021/es048839s
Polak E, Ribiere G (1969) Note sur la convergence de méthodes de directions conjuguées. ESAIM Math Model Numer Anal-Modélisation Mathématique Anal Numér 3:35–43
Poole SK, Poole CF (1999) Chromatographic models for the sorption of neutral organic compounds by soil from water and air. J Chromatogr A 845:381–400. https://doi.org/10.1016/S0021-9673(98)01085-1
Poole CF, Ariyasena TC, Lenca N (2013) Estimation of the environmental properties of compounds from chromatographic measurements and the solvation parameter model. J Chromatogr A 1317:85–104. https://doi.org/10.1016/j.chroma.2013.05.045
Praveen-Kumar, Chhonkar PK, Agnihotri NP (1987) Mobility of urease inhibitors: application of soil thin-layer chromatography. Soil Biol Biochem 19:687–688. https://doi.org/10.1016/0038-0717(87)90049-6
Pussemier L, Szabó G, Bulman RA (1990) Prediction of the soil adsorption coefficient Koc for aromatic pollutants. Chemosphere 21:1199–1212. https://doi.org/10.1016/0045-6535(90)90140-O
Ravanel P, Liégeois MH, Chevallier D, Tissut M (1999) Soil thin-layer chromatography and pesticide mobility through soil microstructures: New technical approach. J Chromatogr A 864:145–154. https://doi.org/10.1016/S0021-9673(99)01007-9
Sobańska AW (2021) RP-18 TLC retention data and calculated physico-chemical parameters as predictors of soil-water partition and bioconcentration of organic sunscreens. Chemosphere 279:130527. https://doi.org/10.1016/j.chemosphere.2021.130527
Sobanska AW, Brzezinska E (2017) Phospholipid-based immobilized artificial membrane (IAM) chromatography: a powerful tool to model drug distribution processes. Curr Pharm Des 23:6784–6704. https://doi.org/10.2174/1381612823666171018114331
Sobańska AW, Robertson J, Brzezińska E (2021) Application of RP-18 TLC retention data to the prediction of the transdermal absorption of drugs. Pharmaceuticals 14:147. https://doi.org/10.3390/ph14020147
Soczewiński E, Wachtmeister CA (1962) The relation between the composition of certain ternary two-phase solvent systems and RM values. J Chromatogr A 7:311–320. https://doi.org/10.1016/s0021-9673(01)86422-0
Sprunger L, Blake-Taylor BH, Wairegi A et al (2007) Characterization of the retention behavior of organic and pharmaceutical drug molecules on an immobilized artificial membrane column with the Abraham model. J Chromatogr A 1160:235–245. https://doi.org/10.1016/j.chroma.2007.05.051
Stergiopoulos C, Makarouni D, Tsantili-Kakoulidou A et al (2019) Immobilized artificial membrane chromatography as a tool for the prediction of ecotoxicity of pesticides. Chemosphere 224:128–139. https://doi.org/10.1016/j.chemosphere.2019.02.075
Szabo G, Prosser SL, Bulman RA (1990a) Prediction of the adsorption coefficient (Koc) for soil by a chemically immobilized humic acid column using RP-HPLC. Chemosphere 21:729–739. https://doi.org/10.1016/0045-6535(90)90260-Z
Szabo G, Prosser SL, Bulman RA (1990b) Determination of the adsorption coefficient (KOC) of some aromatics for soil by RP-HPLC on two immobilized humic acid phases. Chemosphere 21:777–788. https://doi.org/10.1016/0045-6535(90)90265-U
Szabó G, Guczi J, Kördel W et al (1999) Comparison of different HPLC stationary phases for determination of soil-water distribution coefficient, KOC values of organic chemicals in RP-HPLC system. Chemosphere 39:431–442. https://doi.org/10.1016/S0045-6535(99)00006-5
Tao S, Lu X, Cao J, Dawson R (2001) A comparison of the fragment constant and molecular connectivity indices models for normalized sorption coefficient estimation. Water Environ Res 73:307–313. https://doi.org/10.2175/106143001X139326
Tsopelas F, Stergiopoulos C, Tsakanika L-A et al (2017) The use of immobilized artificial membrane chromatography to predict bioconcentration of pharmaceutical compounds. Ecotoxicol Environ Saf 139:150–157. https://doi.org/10.1016/j.ecoenv.2017.01.028
Tsopelas F, Stergiopoulos C, Tsantili-Kakoulidou A (2018) Immobilized artificial membrane chromatography: from medicinal chemistry to environmental sciences. ADMET DMPK 6:225–241. https://doi.org/10.5599/admet.553
US EPA O (2015) EPI SuiteTM-Estimation Program Interface v. 4.11 | US EPA. In: US EPA. https://www.epa.gov/tsca-screening-tools/epi-suitetm-estimation-program-interface. Accessed 22 Aug 2022.
Valko K (2019) Application of biomimetic HPLC to estimate in vivo behavior of early drug discovery compounds. Future Drug Discov 1:FDD11. https://doi.org/10.4155/fdd-2019-0004
Vowles PD, Mantoura RFC (1987) Sediment-water partition coefficients and HPLC retention factors of aromatic hydrocarbons. Chemosphere 16:109–116. https://doi.org/10.1016/0045-6535(87)90114-7
Xu F, Liang X, Lin B et al (1999) Soil column chromatography for correlation between capacity factors and soil organic partition coefficients for eight pesticides. Chemosphere 39:2239–2248. https://doi.org/10.1016/S0045-6535(99)00147-2
Xu F, Liang X, Lin B et al (2002) Estimation of soil organic partition coefficients: from retention factors measured by soil column chromatography with water as eluent. J Chromatogr A 968:7–16. https://doi.org/10.1016/S0021-9673(02)00821-X
Funding
This research was supported by an internal grant of the Medical University of Łódź no. 503/3-016-03/503-31-001.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethical approval
Not applicable
Studies in humans and animals
No human subjects or animal experiments were involved during the preparation of this manuscript.
Consent to participate
Not applicable
Consent for publication
Not applicable
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Marcus Schulz
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sobańska, A.W. Immobilized artificial membrane-chromatographic and computational descriptors in studies of soil-water partition of environmentally relevant compounds. Environ Sci Pollut Res 30, 6192–6200 (2023). https://doi.org/10.1007/s11356-022-22514-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-22514-x