Abstract
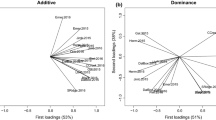
Multi-environment trials (MET) are fundamental for assessing genotype-by-environment interaction (GxE) effects, adaptability and stability of genotypes and provide valuable information about target regions. As such, a MET involving grain sorghum hybrid combinations derived from elite inbred lines adapted to diverse sorghum production regions was developed to assess agronomic performance, stability, and genomic-enabled prediction accuracies within mega-environments (ME). Ten females and ten males from the Texas A&M and Kansas State sorghum breeding programs were crossed following a factorial mating scheme to generate 100 hybrids. Grain yield, plant height, and days to anthesis were assessed in a MET consisting of ten environments across Texas and Kansas over two years. Genotype plus Genotype-by-block-of-environment biplot (GGB) assessed ME, while the "mean-vs-stability" view of the biplot and the Bayesian Finlay–Wilkinson regression evaluated hybrid adaptability and stability. A genomic prediction model including the GxE effect was applied within ME to assess prediction accuracy. Results suggest that grain sorghum hybrid combinations involving lines adapted to different target regions can produce superior hybrids. GGB confirmed distinct regions of sorghum adaption in the U.S. Further, genomic predictions within ME reported inconsistent results, suggesting that additional effects rather than the correlations between environments are influencing genomic prediction of grain sorghum hybrids.







Similar content being viewed by others
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Acosta-Pech R, Crossade los Campos G, J et al (2017) Genomic models with genotype × environment interaction for predicting hybrid performance: an application in maize hybrids. Theor Appl Genet 130:1431–1440. https://doi.org/10.1007/s00122-017-2898-0
Alves FC, Granato ÍSC, Galli G et al (2019) Bayesian analysis and prediction of hybrid performance. Plant Methods 15:14. https://doi.org/10.1186/s13007-019-0388-x
Alves FC, Galli G, Matias FI et al (2021) Impact of the complexity of genotype by environment and dominance modeling on the predictive accuracy of maize hybrids in multi-environment prediction models. Euphytica. https://doi.org/10.1007/s10681-021-02779-y
Ansarifard I, Mostafavi K, Khosroshahli M et al (2020) A study on genotype–environment interaction based on GGE biplot graphical method in sunflower genotypes (Helianthus annuus L.). Food Sci Nutr 8:3327–3334. https://doi.org/10.1002/fsn3.1610
Balestre M, dos Santos VB, Soares AA, Reis MS (2010) Stability and adaptability of upland rice genotypes. Crop Breed Appl Biotechnol 10:357–363. https://doi.org/10.1590/S1984-70332010000400011
Basnet BR, Crossa J, Dreisigacker S et al (2019) Hybrid wheat prediction using genomic, pedigree, and environmental covariables interaction models. Plant Genome 12:1–13. https://doi.org/10.3835/plantgenome2018.07.0051
Bates D, Mächler M, Bolker BM, Walker SC (2015) Fitting linear mixed-effects models using lme4. J Stat Softw. https://doi.org/10.18637/jss.v067.i01
Becker HC (1981) Correlations among some statistical measures of phenotypic stability. Euphytica 30:835–840. https://doi.org/10.1007/BF00038812
Becker HC, Leon J (1988) Stability analysis in plant breeding. Plant Breed 101:1–23. https://doi.org/10.1111/j.1439-0523.1988.tb00261.x
Buntaran H, Forkman J, Piepho H-P (2021) Projecting results of zoned multi-environment trials to new locations using environmental covariates with random coefficient models: accuracy and precision. Theor Appl Genet 1:3. https://doi.org/10.1007/s00122-021-03786-2
Burgueño J, Crossa J, Cornelius PL, Yang RC (2008) Using factor analytic models for joining environments and genotypes without crossover genotype x environment interaction. Crop Sci 48:1291–1305. https://doi.org/10.2135/cropsci2007.11.0632
Burgueño J, de los Campos G, Weigel K, Crossa J (2012) Genomic prediction of breeding values when modeling genotype × environment interaction using pedigree and dense molecular markers. Crop Sci 52:707–719. https://doi.org/10.2135/cropsci2011.06.0299
Coe MT, Evans KM, Gasic K, Main D (2020) Plant breeding capacity in U.S. public institutions. Crop Sci 60:2373–2385. https://doi.org/10.1002/csc2.20227
Comstock RE, Robinson HF (1952) Estimation of average dominance of genes. Heterosis 2:494–516
Cooper M, Messina CD, Podlich D et al (2014) Predicting the future of plant breeding: complementing empirical evaluation with genetic prediction. Crop Pasture Sci 65:311–336. https://doi.org/10.1071/CP14007
Costa-Neto G, Fritsche-Neto R, Crossa J (2020) Nonlinear kernels, dominance, and envirotyping data increase the accuracy of genome-based prediction in multi-environment trials. Heredity (edinb). https://doi.org/10.1038/s41437-020-00353-1
Crossa J, Yang R-C, Cornelius PL (2004) Studying crossover genotype × environment interaction using linear-bilinear models and mixed models. J Agric Biol Environ Stat 9:362–380. https://doi.org/10.1198/108571104X4423
Crossa J, de los Campos G, Pérez P et al (2010) Prediction of genetic values of quantitative traits in plant breeding using pedigree and molecular markers. Genetics 186:713–724. https://doi.org/10.1534/genetics.110.118521
Crossa J, Martini JWR, Gianola D et al (2019) Deep kernel and deep learning for genome-based prediction of single traits in multienvironment breeding trials. Front Genet 10:1–13. https://doi.org/10.3389/fgene.2019.01168
Crossa J, Fritsche-Neto R, Montesinos-Lopez OA et al (2021) The modern plant breeding triangle: optimizing the use of genomics, phenomics, and enviromics data. Front Plant Sci 12:1–6. https://doi.org/10.3389/fpls.2021.651480
Cuevas J, Crossa J, Soberanis V et al (2016) Genomic prediction of genotype × environment interaction kernel regression models. Plant Genome 9:1–20. https://doi.org/10.3835/plantgenome2016.03.0024
Dalló SC, Zdziarski AD, Woyann LG et al (2019) Across year and year-by-year GGE biplot analysis to evaluate soybean performance and stability in multi-environment trials. Euphytica 215:113. https://doi.org/10.1007/s10681-019-2438-x
FAOSTAT (2021) Food and Agriculture Organization of the United Nations. FAOSTAT statistical database. Available online at: http://www.fao.org/faostat/en/#data/QC/visualize
Finlay K, Wilkinson G (1963) The analysis of adaptation in a plant-breeding programme. Aust J Agric Res 14:742. https://doi.org/10.1071/AR9630742
Fonseca J, Perumal R, Klein PE et al (2021b) Combining abilities and elite germplasm enhancement across US public sorghum breeding programs. Crop Sci. https://doi.org/10.1002/csc2.20624
Fonseca JMO, Klein PE, Crossa J et al (2021a) Assessing combining abilities, genomic data, and genotype × environment interactions to predict hybrid grain sorghum performance. Plant Genome. https://doi.org/10.1002/tpg2.20127
Gabriel KR (1971) The biplot graphic display of matrices with application to principal component analysis. Biometrika 58:453–467. https://doi.org/10.1093/biomet/58.3.453
Gauch HG (1988) Model selection and validation for yield trials with interaction. Biometrics 44:705. https://doi.org/10.2307/2531585
Gauch HG Jr (2013) A simple protocol for AMMI analysis of yield trials. Crop Sci 53:1860–1869. https://doi.org/10.2135/cropsci2013.04.0241
Gauch HG, Zobel RW (1997) Identifying mega-environments and targeting genotypes. Crop Sci 37:311–326. https://doi.org/10.2135/cropsci1997.0011183X003700020002x
González-Barrios P, Díaz-García L, Gutiérrez L (2019) Mega-environmental design: using genotype × environment interaction to optimize resources for cultivar testing. Crop Sci 59:1899–1915. https://doi.org/10.2135/cropsci2018.11.0692
Henderson CR (1984) Applications of linear models in animal breeding
Hu X (2015) A comprehensive comparison between ANOVA and BLUP to valuate location-specific genotype effects for rape cultivar trials with random locations. Field Crop Res 179:144–149. https://doi.org/10.1016/j.fcr.2015.04.023
Huehn M (1990) Nonparametric measures of phenotypic stability. Part 1: theory. Euphytica 47:189–194. https://doi.org/10.1007/BF00024241
Jarquín D, Crossa J, Lacaze X et al (2014) A reaction norm model for genomic selection using high-dimensional genomic and environmental data. Theor Appl Genet 127:595–607. https://doi.org/10.1007/s00122-013-2243-1
Jarquin D, Howard R, Crossa J et al (2020) Genomic prediction enhanced sparse testing for multi-environment trials. G3 Genes. Genomes Genet 10:2725–2739. https://doi.org/10.1534/g3.120.401349
Kofoid KD, Harvey TL (2005) Registration of greenbug resistant sorghum germplasm lines KS 116 A/B through KS 120 A/B. Crop Sci 45:802. https://doi.org/10.2135/cropsci2005.0802
Kuznetsova A, Brockhoff PB, Christensen RHB (2017) lmerTest package tests in linear mixed effects models. J Stat Softw. https://doi.org/10.18637/jss.v082.i13
Lado B, Barrios PG, Quincke M et al (2016) Modeling genotype × environment interaction for genomic selection with unbalanced data from a wheat breeding program. Crop Sci 56:2165–2179. https://doi.org/10.2135/cropsci2015.04.0207
Laffont JL, Wright K, Hanafi M (2013) Genotype plus genotype × block of environments biplots. Crop Sci 53:2332–2341. https://doi.org/10.2135/cropsci2013.03.0178
Lenth R (2021) emmeans: estimated marginal means, aka least-squares means. R Packag. version 1.6.0
Li X, Guo T, Mu Q et al (2018) Genomic and environmental determinants and their interplay underlying phenotypic plasticity. Proc Natl Acad Sci U S A 115:6679–6684. https://doi.org/10.1073/pnas.1718326115
Lian L, de los Campos G (2016) FW an R package for finlay-Wilkinson regression that incorporates genomic/pedigree information and covariance structures between environments. G3 Genes Genom Genet 6:589–597. https://doi.org/10.1534/g3.115.026328
Lopez-Cruz M, Crossa J, Bonnett D et al (2015) Increased prediction accuracy in wheat breeding trials using a marker × environment interaction genomic selection model. G3 Genes Genom Genet 5:569–582. https://doi.org/10.1534/g3.114.016097
Mbulwe L, Peterson GC, Scott-Armstrong J, Rooney WL (2015) Registration of sorghum germplasm Tx3408 and Tx3409 with tolerance to sugarcane aphid [Melanaphis sacchari (Zehntner)]. J Plant Regist 10:51–56. https://doi.org/10.3198/jpr2015.04.0025crg
Miller FR, Dusek TF, Prihoda KL, Rooney LW (1992) Registration of RTx436 sorghum parental line. Crop Sci 32:1518–1518. https://doi.org/10.2135/cropsci1992.0011183x003200060059x
Monk R, Franks C, Dahlberg J (2014) Sorghum. In: Yield gains in major US field crops. Wiley Online Library, pp 293–310
NASA 2021. Prediction Of Worldwide Energy Resources (POWER). Available online at: https://power.larc.nasa.gov/data-access-viewer/. Accessed 16 April 2021
NASS (2021a) 2020 state agriculture overview: Texas. USDANational Agricultural Statistics Service. https://www.nass.usda.gov/Quick_Stats/Ag_Overview/stateOverview.php?state=TEXAS. Accessed 25 Mar 2021a
NASS (2021b) 2020 state agriculture overview: Kansas.USDA National Agricultural Statistics Service. https://www.nass.usda.gov/Quick_Stats/Ag_Overview/stateOverview.php?state=KANSAS. Accessed 25 Mar 2021b
NASS (U. S. Department of Agriculture National Agricultural Statistics Service) (2020) Crop production. Available online at: https://www.nass.usda.gov/Publications/Todays_Reports/reports/crop1020.pdf. Accessed 5 Feb 2021
Nassar R, Huhn M (1987) Studies on estimation of phenotypic stability: tests of significance for nonparametric measures of phenotypic stability. Biometrics 43:45. https://doi.org/10.2307/2531947
Nielsen DC, Vigil MF (2018) Wheat yield and yield stability of eight dryland crop rotations. Agron J 110:594–601. https://doi.org/10.2134/agronj2017.07.0407
Patterson HD, Thompson R (1971) Recovery of inter-block information when block sizes are unequal. Biometrika 58:545–554. https://doi.org/10.1093/biomet/58.3.545
Pérezde los Campos G P (2014) Genome-wide regression and prediction with the BGLR statistical package. Genetics 198:483–495. https://doi.org/10.1534/genetics.114.164442
Perumal R, Tesso T, Kofoid KD et al (2019) Registration of six grain sorghum pollinator (R) lines. J Plant Regist 13:113–117. https://doi.org/10.3198/jpr2017.12.0087crp
Piepho HP (1994) Best linear unbiased prediction (BLUP) for regional yield trials: a comparison to additive main effects and multiplicative interaction (AMMI) analysis. Theor Appl Genet 89:647–654. https://doi.org/10.1007/BF00222462
Piepho H-P (1998) Empirical best linear unbiased prediction in cultivar trials using factor-analytic variance-covariance structures. Springer
Piepho HP, Möhring J (2005) Best linear unbiased prediction of cultivar effects for subdivided target regions. Crop Sci 45:1151–1159. https://doi.org/10.2135/cropsci2004.0398
Piepho HP, Möhring J, Melchinger AE, Büchse A (2008) BLUP for phenotypic selection in plant breeding and variety testing. Euphytica 161:209–228
Piepho HP (1997) Mixed models with multiplicative terms for cultivar trial data. Adv Biom Genet 215–220
Podlich DW, Cooper M (1998) Modelling plant breeding programs as search strategies on a complex response surface. In: Asia-Pacific conference on simulated evolution and learning. Springer, pp 171–178
R Core Team (2020) R: a language and environment for statistical computing
Rakshit S, Ganapathy KN, Gomashe SS et al (2012) GGE biplot analysis to evaluate genotype, environment and their interactions in sorghum multi-location data. Euphytica 185:465–479. https://doi.org/10.1007/s10681-012-0648-6
Resende MDV, Thompson R (2004) Factor analytic multiplicative mixed models in the analysis of multiple experiments. Rev Matemática e Estatística São Paulo 22:31–52
Rooney WL (2016) Sorghum: production and improvement practices, 2nd edn. Elsevier
Rooney WL, Miller FR, Rooney LW (2003) Registration of RTx437 sorghum parental line. Crop Sci 43:445. https://doi.org/10.2135/cropsci2003.4450
Rosenow DT, Clark LE, Peterson GC et al (2021) Registration of Tx3440 through Tx3482 sorghum germplasm. J Plant Regist. https://doi.org/10.1002/plr2.20082
Sharma SP, Leskovar DI, Crosby KM, Ibrahim AMH (2020) GGE biplot analysis of genotype-by-environment interactions for melon fruit yield and quality traits. HortScience 55:533–542
Shelton AC, Tracy WF (2017) Cultivar development in the U.S. public sector. Crop Sci 57:1823–1835. https://doi.org/10.2135/cropsci2016.11.0961
Shukla GK (1972) Some statistical aspects of partitioning genotype environmental components of variability. Heredity (edinb) 29:237–245
Smith A, Cullis B, Thompson R (2001) Analyzing variety by environment data using multiplicative mixed models and adjustments for spatial field trend. Biometrics 57:1138–1147. https://doi.org/10.1111/j.0006-341X.2001.01138.x
Smith AB, Cullis BR, Thompson R (2005) The analysis of crop cultivar breeding and evaluation trials: an overview of current mixed model approaches. J Agric Sci 143:449–462
Smith AB, Borg LM, Gogel BJ, Cullis BR (2019) Estimation of factor analytic mixed models for the analysis of multi-treatment multi-environment trial data. J Agric Biol Environ Stat 24:573–588. https://doi.org/10.1007/s13253-019-00362-6
Sorensen D, Gianola D (2007) Likelihood, bayesian, and MCMC methods in quantitative genetics. Springer
Technow F, Schrag TA, Schipprack W et al (2014) Genome properties and prospects of genomic prediction of hybrid performance in a breeding program of maize. Genetics 197:1343–1355. https://doi.org/10.1534/genetics.114.165860
Technow F, Podlich D, Cooper M (2020) Back to the future: implications of genetic complexity for hybrid breeding strategies. BioRxiv. https://doi.org/10.1101/2020.10.21.349332
Wricke G (1962) Uber eine methode zur erfassung der okologischen streubreite in feldverzuchen. Z Pflanzenzuchtg 47:92–96
Wright K, Laffont J-L (2020) gge: genotype plus genotype-by-environment biplots. R Packag. version 1.6
Yan W (2015) Mega-environment analysis and test location evaluation based on unbalanced multiyear data. Crop Sci 55:113–122. https://doi.org/10.2135/cropsci2014.03.0203
Yan W (2016) Analysis and handling of G × E in a practical breeding program. Crop Sci 56:2106–2118
Yan W, Kang MS (2002) GGE biplot analysis: a graphical tool for breeders, geneticists, and agronomists. CRC Press
Yan W, Hunt LA, Sheng Q, Szlavnics Z (2000) Cultivar evaluation and mega-environment investigation based on the GGE biplot. Crop Sci 40:597–605. https://doi.org/10.2135/cropsci2000.403597x
Acknowledgements
Funds from the Borlaug-Monsanto Chair in Plant Breeding were used to support this work. Additionally, the authors wish to thank Dr. Costa-Neto for suggesting the Bayesian Finlay–Wilkinson regression presented herein.
Author information
Authors and Affiliations
Contributions
JMOF: Conceptualization; Data curation; Formal analysis; Funding acquisition; Investigation; Methodology; Visualization; Writing-original draft; Writing-review & editing. PR: Conceptualization; Investigation; Resources; Writing-review & editing. PEK: Investigation; Writing-review & editing. RK: Investigation; Writing-review & editing. WLR: Conceptualization; Funding acquisition; Investigation; Methodology; Project administration; Resources; Supervision; Writing-review & editing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fonseca, J.M.O., Perumal, R., Klein, P.E. et al. Mega-environment analysis to assess adaptability, stability, and genomic predictions in grain sorghum hybrids. Euphytica 218, 128 (2022). https://doi.org/10.1007/s10681-022-03075-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-022-03075-z